Mechanical testing
We test five plant-based meat products: ham style roast tofurky (Tofurky, Hood River, OR), artesian vegan frankfurter plant sausage (Field Roast, Seattle, WA), signature stadium plant hotdog (Field Roast, Seattle, WA), organic firm tofu (365 by Whole Foods Market, Austin TX), organic extrafirm tofu (House Foods, Garden Grove, CA). For comparison, we also tested three animal meat products: wieners classic animal hotdog (Oscar Meyer, Kraft Heinz Co, Chicago, IL), oven roasted spam turkey (Spam, Hormel Foods Co, Austin, MN), and turkey polska kielbasa animal sausage (Hillshire Farm, New London, WI). Table 1 summarizes the ingredients of all eight products. For each meat type, we test at least n = 5 samples in tension, compression, and shear. Figure 6 documents our sample preparation and our mechanical testing.
a Samples are cut using a 3D printed guide and a brain sectioning knife to obtain uniform dimensions of 1 × 1 × 2 cm3 for tensile testing. b Samples are super glued to glass microscope slides. c Samples are left to set for 30-45 minutes for the glue to adhere. d Cylindrical samples of 8 mm diameter and 1 cm height are cored using a biopsy punch and loaded into a rheometer for compression and shear testing using parallel plates with sandpaper on both surfaces. e Tension samples are loaded into 3D printed grips, which are compatible with an Instron testing device. f Tension test is run until the sample fails. a, b, d show animal sausage and c, e, f show animal hotdog.
Sample preparation
We prepare the samples following our established protocols28. Figure 6b, c, e, f illustrate our tension tests, for which we use a custom 3D printed cutting guide and brain sectioning knife to prepare samples of 1 × 1 × 2 cm3. We super glue the samples to glass microscope slides and wait for 30 min until the glue is fully cured. During curing, we drape a damp paper towel over the samples to keep them hydrated. For the compression and shear tests, we prepare cylindrical samples of 8 mm diameter and 1 cm height using a biopsy punch to extract full-thickness cores from the center of each material, and store the samples in a damp paper tower until testing.
Sample testing
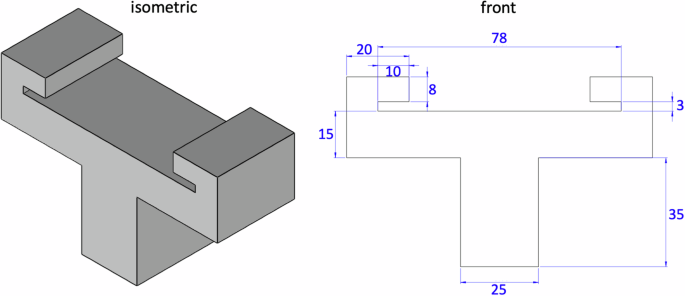
For all three modes, tension, compression, and shear, we test the samples raw and at room temperature at 25 °C28. We perform all uniaxial tension tests using an Instron 5848 (Instron, Canton, MA) with a 100N load cell, see Fig. 6e. We use 3D printed custom grips to rapidly mount and unmount the microscope slides for high throughput testing. Figure 7 shows the part dimensions to create these grips. We mount each sample in the grips, apply a small pre-load, and calibrate the initial gauge length L. We determine the pre-load magnitude for each sample individually. We define pre-load as the minimum load needed to remove any slack, based on visual inspection of a force-displacement curve starting with a fully unloaded specimen, generally on the order of 0.5 N. We then increase the tensile stretch quasi-statically at a rate of (dot{lambda }=0.2 %)/s for t = 50 s, until the sample fails. We perform all uniaxial compression and shear tests using an AR-2000ex torsional rheometer (TA Instruments, New Castle, DE), see Fig. 6d. For the compression tests, we mount the sample, apply a small pre-load, and calibrate the initial gage length L. We determine the pre-load for each sample based on its loading curve, with values on the order of 0.5 N. We then increase the compressive stretch quasi-statically at a rate of (dot{lambda }=0.2 %)/s for t = 50 s, up to a total stretch of λ = 0.9. For the shear tests, we apply a 10% compressive pre-load and calibrate the initial gage length L. We then rotate the upper plate quasi-statically at a shear rate of (dot{gamma }=0.2 %)/s for t = 50 s, up to a total shear of γ = 0.1. To prevent slippage of the samples during the shear tests, we use a sandpaper-covered base plate of 20 mm diameter and a sandpaper-covered top plate of 8 mm diameter.
Front, right, and isometric views of our customized 3D printed device to hold glass slides during tensile testing. Two holders need to be printed to complete the set-up. A standard microscope glass slide fits into the 3 mm slot; it mounts and unmounts easily for high throughout testing. All dimensions are in mm.
Analytical methods and data processing
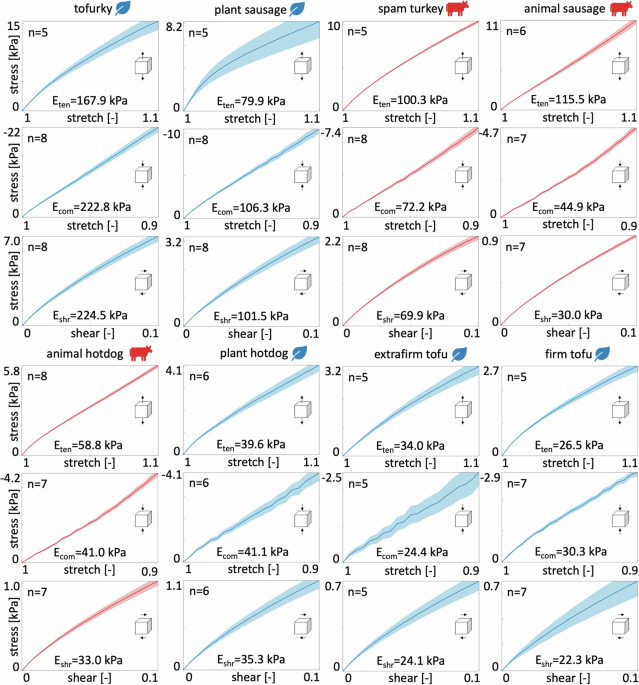
For each sample and each test mode, we use MATLAB (Mathworks, Natick, MA, USA) to smooth the curves using smoothingspline and SmoothingParam = 1. We interpolate each smooth curve over 21 equidistant points in the ranges (1.0le lambda le {lambda }_{max }) for tension, 1.0 ≥ λ ≥ 0.9 for compression, and 0.0 ≤ γ ≤ 1.0 for shear. For each meat product, we select ({lambda }_{max }) in tension as the maximum stretch in the hyperelastic regime, the loading range within which we observe no visible failure for any of the samples. Finally, we average the interpolated curves to obtain the mean and standard error of the mean for each product and report the data in Table 2.
Stiffness
For each testing mode, we extract the stiffness of each product from the data in Table 2 using linear regression. We convert that the tension and compression data {λ; P11} into strain-stress pairs {ε; σ}, where ε = λ − 1 is the strain and σ = P11 is the stress. We postulate a linear stress-strain relation, σ = E ⋅ ε, and use linear regression to determine the tensile and compressive stiffnesses ({E}_{{{rm{ten}}}},{E}_{{{rm{com}}}}=({{boldsymbol{varepsilon }}}cdot hat{{{boldsymbol{sigma }}}})/({{boldsymbol{varepsilon }}}cdot {{boldsymbol{varepsilon }}})). Similarly, we rewrite the shear data, {γ; P12}, as shear strain-stress pairs {γ; τ}, where γ is the shear strain and τ = P12 is the shear stress. We postulate a linear shear stress-strain relation, τ = μ ⋅ γ, convert the shear modulus μ into the shear stiffness, Eshr = 2 [ 1 + ν ] μ = 3 μ, and use linear regression, to determine the shear stiffness ({E}_{{{rm{shr}}}}=3,({{boldsymbol{gamma }}}cdot hat{{{boldsymbol{tau }}}})/({{boldsymbol{gamma }}}cdot {{boldsymbol{gamma }}})).
Kinematics
We analyze all three testing modes combined using finite deformation continuum mechanics42,43. During testing, particles X of the undeformed sample map to particles, x = φ(X), of the deformed sample via the deformation map φ. Similarly, line elements of the dX of the undeformed sample map to line elements, dx = F ⋅ dX, of the deformed sample via the deformation gradient, ({{boldsymbol{F}}}={nabla }_{{{boldsymbol{X}}}}{{boldsymbol{varphi }}}={sum }_{i = 1}^{3},{lambda }_{i},{{{boldsymbol{n}}}}_{i}otimes {{{boldsymbol{N}}}}_{i}). Its spectral representation introduces the principal stretches λi and the principal directions Ni and ni in the undeformed and deformed configurations, where F ⋅ Ni = λini. We assume that all meat samples are isotropic and have three principal invariants, ({I}_{1}={lambda }_{1}^{2}+{lambda }_{2}^{2}+{lambda }_{3}^{2}) and ({I}_{2}={lambda }_{1}^{2}{lambda }_{2}^{2}+{lambda }_{2}^{2}{lambda }_{3}^{2}+{lambda }_{1}^{2}{lambda }_{3}^{2}) and ({I}_{3}={lambda }_{1}^{2},{lambda }_{2}^{2},{lambda }_{3}^{2}={J}^{2}), which are linear, quadratic, and cubic in terms of the principal stretches squared. We also assume that all samples are perfectly incompressible, and their third invariant always remains equal to one, I3 = 1. The remaining two invariants, I1 and I2, depend on the type of experiment.
Constitutive equations
Constitutive equations relate a stress like the Piola or nominal stress P, the force per undeformed area that we measure during our experiments, to a deformation measure like the deformation gradient F28. For a hyperelastic material that satisfies the second law of thermodynamics, we can express the Piola stress, P = ∂ψ(F)/∂F − p F−t, as the derivative of the Helmholtz free energy function ψ(F) with respect to the deformation gradient F, modified by a pressure term, − p F−t, to ensure perfect incompressibility. Here, the hydrostatic pressure, (p=-frac{1}{3},{{boldsymbol{P}}}:{{boldsymbol{F}}}), acts as a Lagrange multiplier that that we determine from the boundary conditions of our experiments. Instead of formulating the free energy function directly in terms of the deformation gradient ψ(F), we can express it in terms of the invariants, ψ(I1, I2), and obtain the following expression, P = ∂ψ/∂I1 ⋅ ∂I1/∂F + ∂ψ/∂I2 ⋅ ∂I2/∂F − p F−t.
Constitutive neural networks
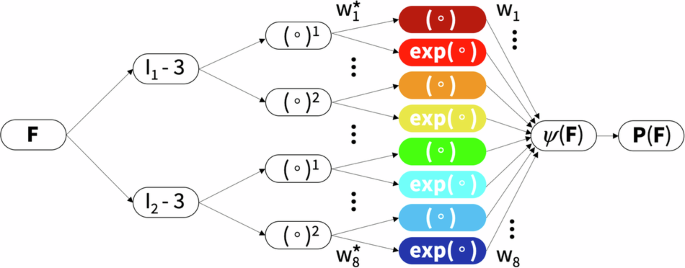
Motivated by these kinematic and constitutive considerations, we reverse-engineer our own constitutive neural network that satisfies the conditions of thermodynamic consistency, material objectivity, material symmetry, incompressibility, constitutive restrictions, and polyconvexity by design29,44. Yet, instead of building these constraints into the loss function, we hardwire them directly into our network input, output, architecture, and activation functions45,46 to satisfy the fundamental laws of physics. Special members of this family represent well-known constitutive models, including the neo Hooke32, Blatz Ko30, Mooney Rivlin33,34, and Demiray31 models, for which the network weights gain a clear physical interpretation44,47. Specifically, our constitutive neural network learns a free energy function that is parameterized in terms of the first and second invariants. It takes the deformation gradient F as input, computes the two invariants, I1 and I2, raises them to the first and second powers, (∘) and (∘)2, applies either the identity or exponential function, (∘) and exp(∘), and summarizes all eight terms in the strain energy function ψ as the network output. Figure 8 illustrates our network with n = 8 nodes, with the following eight-term free energy function,
$$begin{array}{lllllll}psi(I_1, I_2)& = & w_1 &[I_1-3] &+& w_2 & [exp(w_2^* [I_1-3])-1] \ & + & w_3 &[I_1-3]^2 &+& w_4 & [exp(w_4^* [I_1-3]^2) -1] \ & + & w_5 &[I_2-3] &+& w_6 & [exp(w_6^* [I_2-3])-1] \ & + & w_7 &[I_2-3]^2 &+& w_8 & [exp(w_8^* [I_2-3]^2) -1] , ,end{array}$$
where w = [ w1, w2, w3, w4, w5, w6, w7, w8 ] and ({{{boldsymbol{w}}}}^{* }=[,{w}_{2}^{* },{w}_{4}^{* },{w}_{6}^{* },{w}_{8}^{* }]) are the network weights. From the derivative of the free energy, we calculate the stress,
$$begin{array}{lllll}{boldsymbol{P}}&=&&& left[right. w_1+w_2 exp(w_2^* [I_1-3]) \ &+&2&[I_1-3]&[w_3 +w_4 exp(w_4^*[I_1-3]^2)]left.right]partial I_1/partial boldsymbol{F}\ &+&&&left[right. w_5+w_6 exp(w_6^* [I_2-3]) \ &+&2&[I_2-3]&[w_7 +w_8 exp(w_8^*[I_2-3]^2)]left.right]partial I_2/partial boldsymbol{F}, end{array}$$
where the derivatives of the invariants, ∂I1/∂F and ∂I2/∂F, depend on the type or experiment. During training, our network autonomously discovers the best subset of activation functions from 28 − 1 = 255 possible combinations of terms. At the same time, it naturally trains the weights of the less important terms to zero.
Isotropic, perfectly incompressible constitutive artificial neural network with two hidden layers and eight terms. The network takes the deformation gradient F as input and calculates its first and second invariant terms, [I1 − 3] and [I2 − 3]. The first layer generates powers of these invariants, (∘)1 and (∘)2, and the second layer applies the identity and the exponential function to these powers, (∘) and exp(∘). The strain energy function ψ(F) is a sum of the resulting eight terms. Its derivative defines the Piola stress, ∂ψ(F)/∂F, whose components, P11 or P12, enter the loss function to minimize the error with respect to the tension, compression, and shear data. By minimizing the loss function, the network trains its weights w and w* and discovers the best model and parameters to explain the experimental data.
Uniaxial tension and compression
In the tension and compression experiments, we apply a stretch λ = l/L, that we calculate as the ratio between the current and initial sample lengths l and L. We can write the deformation gradient F in matrix representation as
$${{boldsymbol{F}}}=left[begin{array}{ccc}lambda &0&0\ 0&1/sqrt{lambda }&0\ 0&0&1/sqrt{lambda }end{array}right]quad {{rm{with}}}quad lambda =l/L.$$
In tension and compression, the first and second invariants and their derivatives are I1 = λ2 + 2/λ and I2 = 2λ + 1/λ2 with ∂λI1 = 2 λ − 2/λ2 and ∂λI2 = 2 − 2/λ3. Using the zero normal stress condition, P22 = P33 = 0, we obtain the explicit expression for the uniaxial stress, P11 = 2 [λ − 1/λ2] ∂ψ/∂I1 + 2 [1 − 1/λ3] ∂ψ/∂I2, which we can write explicitly in terms of the network weights w and w*,
$$begin{array}{lllll}P_{11}&=&2&left[lambda-{1}/{lambda^2}right]&left[,w_1+w_2 exp(w_2^*[I_1-3])right.\ &+&2&[I_1-3]&[,w_3 +w_4 exp(w_4^*[I_1-3]^2)]left.right]\ &+&2&left[1-{1}/{lambda^3}right]&left[right.,w_5+w_6 exp(w_6^*[I_2-3])\ &+&2&[I_2-3]&[,w_7 +w_8 exp(w_8^*[I_2-3]^2)]left.right].end{array}$$
Simple shear
In the shear experiment, we apply a torsion angle ϕ, that translates into the shear stress, γ = r/L ϕ, by multiplying it with the sample radius r and dividing by the initial sample length L. We can write the deformation gradient F in matrix representation as
$${{boldsymbol{F}}}=left[begin{array}{ccc}1&gamma &0\ 0&1&0\ 0&0&1end{array}right]quad {{rm{with}}}quad gamma =r/L,phi .$$
In shear, the first and second invariants and their derivatives are I1 = 3 + γ2 and I2 = 3 + γ2 with ∂γI1 = 2 γ and ∂γI2 = 2 γ. We obtain the explicit expression for the shear stress, P12 = 2 [∂ψ/∂I1 + ∂ψ/∂I2] γ, which we can write explicitly in terms of the network weights w and w*,
$$begin{array}{lllll}P_{12}&=&2&gamma&left[,w_1+w_2 exp(w_2^*[I_1-3])right.\ &+&2&[I_1-3]&left.[,w_3 +w_4 exp(w_4^*[I_1-3]^2)]right]\ &+&2&gamma&left[,w_5+w_6 exp(w_6^*[I_2-3])right.\ &+&2&[I_2-3]&left.[,w_7 +w_8 exp(w_8^*[I_2-3]^2)]right]. end{array}$$
Loss function
Our constitutive neural network learns the network weights, w and w*, by minimizing the loss function L that penalizes the mean squared error, the L2-norm of the difference between model and data, divided by the number of data points in tension, compression, and shear28,
$$begin{array}{lllll}L& = &frac{1}{n_{rm{ten}}}&sumlimits_{i=1}^{n_{rm{ten}}} & |,, P_{rm{ten}} (lambda_i) – hat{P}_{rm{ten},i},, ||^2 \ & + &frac{1}{n_{rm{com}}}&sumlimits_{i=1}^{n_{rm{com}}} & |,, P_{rm{com}} (lambda_i) – hat{P}_{rm{com},i},, |^2 \ & + &frac{1}{n_{rm{shr}}}&sumlimits_{i=1}^{n_{rm{shr}}} & | ,,P_{rm{shr}} (gamma_i ) – hat{P}_{rm{shr},i},, |^2 rightarrow minend{array}$$
We train the network by minimizing the loss function with the ADAM optimizer, a robust adaptive algorithm for gradient-based first-order optimization using the tension, compression, and shear data for all eight meat products from Table 2.
Best-in-class modeling
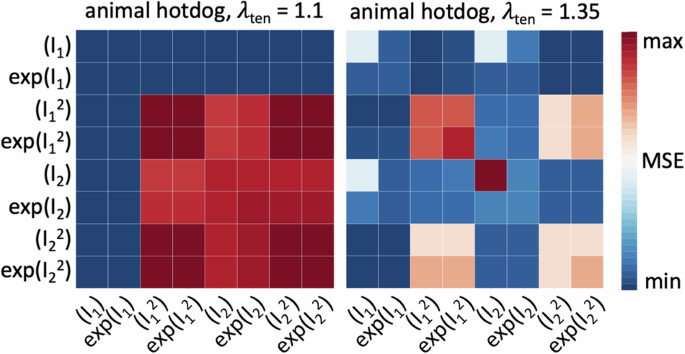
Instead of looking for the best possible fit of the models to the experimental data, we seek to discover meaningful constitutive models that are interpretable and generalizable48, models that have a sparse parameter vector with a limited number of terms49. From combinatorics, we know that our network can discover 28 − 1 = 255 possible models, 8 with a single term, 28 with two, 56 with three, 70 with four, 56 with five, 28 with six, 8 with seven, and 1 with all eight terms. For practical purposes, we focus on the the 8 one-term models and the 28 two-term models, set all other weights to zero, and discover the non-zero weights that characterize the active terms. We summarize the results in a color-coded 8 × 8 error plot, as the average of the mean squared error across the tension, compression, and shear data, and report the parameters of the best-in-class one- and two-term models in Table 1. Motivated by the notable differences in the discovered models for tensile stretches up to 10% versus 35% in Fig. 9, we decide to use the full tension data, not just the 10% stretch.
Discovered one-term models, on the diagonal, and two-term models, off-diagonal, using tensile stretches up to 10% versus up to the peak stress of 35%. All models are made up of eight functional building blocks: linear, exponential linear, quadratic, and exponential quadratic terms of the first and second strain invariants I1 and I2. The color code indicates the quality of fit to the tension, compression, and shear data from Table 2, ranging from dark blue, best fit, to dark red, worst fit. The larger stretch range of 35% provides a clearer distinction of the quality of fit for the individual models.
Food texture survey
We prepare bite-sized samples of the eight products, five plant-based, tofurky, plant-based sausage, plant-based hotdog, extrafirm tofu, and firm tofu, and three animal-based, spam turkey, animal sausage, and animal hotdog. We bake all samples in an oven until the plant-based products sre sufficiently warm, and the animal products reach a safe internal temperature for eating. We do not add any sauces or condiments, except for a small bit of oil to prevent excess sticking to the parchment paper. We keep the samples warm until serving. We recruit n = 16 participants to participate in three surveys: the ten-question Food Neophobia Survey35 and the sixteen-question Meat Attachment Questionnaire36, see Fig. 10, and our own Food Texture Survey. We instruct each participant to eat a sample of each meat product and rank its texture features according to our survey. The survey uses a 5-point Likert scale with twelve questions. Each question starts with “this food is …”, followed by one of the following features21,22: soft, hard, brittle, chewy, gummy, viscous, springy, sticky, fibrous, fatty, moist, and meat-like. The scale ranges from 5 for strongly agree to 1 for strongly disagree. This research was reviewed and approved by the Institutional Review Board at Stanford University under the protocol IRB-75418.
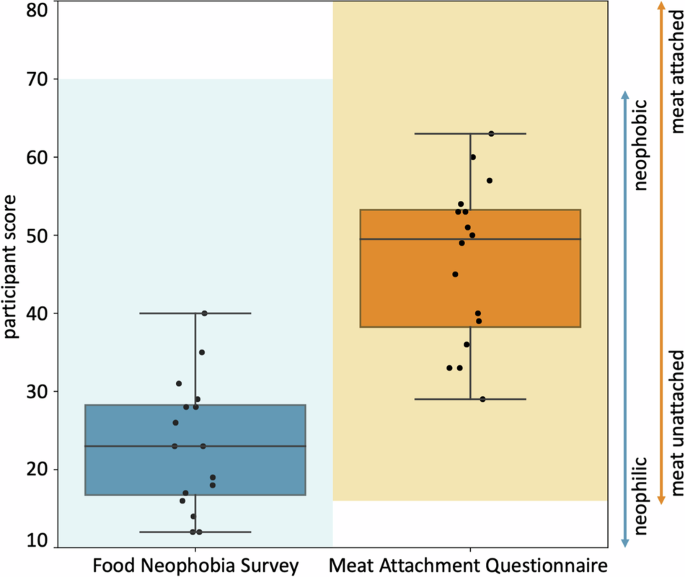
The Food Neophobia Survey uses a 7-point Likert scale with 10 questions so the light blue shaded range goes from 10 (neophilic, open to trying new foods) to 70 (neophobic, not open to trying new foods)35. The Meat Attachment Questionnaire uses a 5-point Likert scale with 16 questions so the light orange range goes from 16 (unattached to eating meat) to 80 (very attached to eating meat)36. The box-and-whisker plots of the minimum, first quartile, median, third quartile, and maximum participant scores are plotted in dark blue and dark orange for the Food Neophobia Survey and Meat Attachment Questionnaire, respectively. The black dots show individual scores.